A tour of probabilistic programming language APIs
What does it look like to do MCMC in different frameworks?
Introduction
I wanted an easy reference for myself and others to see how different developers think about defining
probabilistic models, and this is an attempt at that. I have a number of biases
I am a contributor to PyMC3, and have been working on
PyMC4 (which uses TensorFlow probability). I write far more Python than R, and far more
R than julia or C++. , but the code
shown here is open-source, and contributions/suggestions are gratefully accepted.
I am also including an opinionated - but I hope fair - summary of the projects as of the time of
writing.
On my TODO list are greta in R;
Turing and Gen in
Julia; and Figaro and Rainier in Scala.
Different libraries are written for different reasons, and I care strongly about easily writing and
sharing models, and then sampling from them.
The Task
I have generated a dataset as follows:
import numpy as np np.random.seed(0) ndims = 5 ndata = 100 X = np.random.randn(ndata, ndims) w_ = np.random.randn(ndims) # hidden noise_ = 0.1 * np.random.randn(ndata) # hidden y_obs = X.dot(w_) + noise_
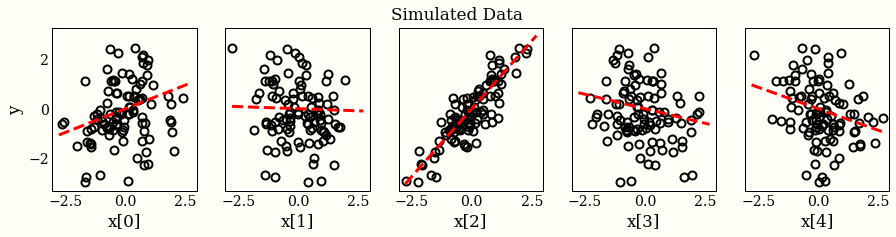
For each probabilistic programming language (PPL), I will:
- Write down the following model: $$ \begin{align} p(\mathbf{w}) &\sim \mathcal{N}(\mathbf{0}, I_5)\\ p(\mathbf{y} | X, \mathbf{w}) &\sim \mathcal{N}(X\mathbf{w}, 0.1I_{100}), \end{align} $$ where \(I_n\) is the \(n \times n\) identity matrix.
- Draw 1,000 samples from the posterior distribution
$$
p(\mathbf{w} | X, \mathbf{y}) \propto p(\mathbf{y} | X, \mathbf{w}) p(\mathbf{w})
$$
where I pass
y_obsto the code at some point. - Do a sanity check that the samples for
ware reasonable near to the true values forw, given the model.For those keeping track at home, the posterior of the model given the data is also a Gaussian, $$p(\mathbf{w} | X, \mathbf{y}) = \mathcal{N}((X^TX + I_5)^{-1}X^T\mathbf{y}, (X^TX + I_5)^{-1})$$
PyStan
version 2.19.0.0Documentation for PyStan, and for Stan itself.
Hand-rolled automatic differentiation in C++. Many (most?) good ideas in PyMC3 came from
here. Updates to the code sometimes accompany papers describing the new algorithms. The
authors/contributors care deeply about correctness and practicality, and it shows. A very good, fast
choice if you care about MCMC. The models are in their own programming language, which can be a little
funny to use in Python (where I would prefer Python objects to strings). Uses the dynamic NUTS
algorithmThe Stan team actually keeps
changing/improving the sampling algorithm they use, and calling it NUTS is at least a little
misleading, since these improvements are substantive compared to the algorithm described in
the paper. If you want to be proper, call it something like "dynamic Hamiltonian Monte Carlo method
with multinomial sampling of dynamic length trajectories, generalized termination criterion, and
improved adaptation of the Euclidean metric." by default, which is currently the best default
to use, though only two libraries here have a good version.
import pystan
linear_regression = """
data {
int N; // number of data items
int K; // number of predictors
matrix[N, K] X; // predictor matrix
vector[N] y; // outcome vector
}
parameters {
vector[K] w; // coefficients for predictors
}
model {
y ~ normal(X * w, 0.1); // likelihood
}
"""
linear_data = {'N': ndata,
'K': ndims,
'y': y_obs,
'X': X}
sm = pystan.StanModel(model_code=linear_regression)
fit = sm.sampling(data=linear_data, iter=1000, chains=4)
PyMC3
version 3.7Uses theano for automatic differentiation. The contributors are strong, all the diagnostics
are good looking, and all the estimates are above average See my biases above.. More seriously, it is
quite fast, flexible if you are willing to write Theano, and has robust tuning and diagnostics, so you
can get some samples from a model with a minimum of fuss. Interoperates quite smoothly with Python
projects. Also uses the NUTS algorithm.
import pymc3 as pm
import theano.tensor as tt
with pm.Model():
w = pm.Normal('w', 0, 1, shape=ndims)
y = pm.Normal('y', tt.dot(X, w), 0.1, observed=y_obs)
trace = pm.sample(1000)
emcee
version 2.2.1This library is a "pure-Python implementation of Goodman & Weare’s Affine Invariant Markov chain Monte Carlo (MCMC) Ensemble sampler", and does not use gradients. This means it does not scale as well to over, say 10 dimensions, but installation is very easy. Also somewhat unique in writing custom likelihood and prior density functions. Lots of examples and users in the astrophysics community.
import scipy.stats as st
import emcee
# log likelihood
def lnlike(w, X, y):
model = X.dot(w)
inv_sigma2 = 0.1 ** -2
return -0.5*(np.sum((y-model)**2)*inv_sigma2 - np.log(inv_sigma2))
# Define a prior for w
w_rv = st.multivariate_normal(np.zeros(ndims), np.eye(ndims))
# Log probability for w
lnprior = w_rv.logpdf
# logp(w | X, y) = logp(y | X, w) + logp(w)
def lnprob(w, X, y):
return lnprior(w) + lnlike(w, X, y)
nwalkers = 100
pos = w_rv.rvs(size=nwalkers)
sampler = emcee.EnsembleSampler(nwalkers, ndims, lnprob, args=(X, y_obs))
pos, lprob, rstate = sampler.run_mcmc(pos, 1000)
Pyro
version 0.3.4Uses pytorch for automatic differentiation. I get the impression the library traditionally
cared more about variational inference, but running HMCAs is the case with TensorFlow Probability, this
implementation is parametrized by number of steps rather than path length, which has some
performance implications I still do not entirely grasp. For example, taking smaller steps with a
fixed path length means each sample takes longer, while keeping a fixed number of steps means the
sample will take the same amount of time but be more correlated. was quite smooth.
There is no NUTS, which means you may have to manually set the number of stepsThis was clearly documented, I just missed it. Updated the code
example to use it, too.. Pyro implements NUTS as well as HMC.
import pyro
import torch
from pyro.infer.mcmc import NUTS, MCMC
import pyro.distributions as dist
def model(X):
w = pyro.sample('w', dist.Normal(torch.zeros(ndims), torch.ones(ndims)))
y = pyro.sample('y', dist.Normal(torch.matmul(X, w), 0.1 * torch.ones(ndata)),
obs=torch.as_tensor(y_obs, dtype=torch.float32))
return y
nuts_kernel = NUTS(model, adapt_step_size=True)
py_mcmc = MCMC(nuts_kernel, num_samples=1_000, warmup_steps=500)
py_mcmc = py_mcmc.run(torch.as_tensor(X, dtype=torch.float32))
TensorFlow Probability
version 0.8.0-dev20190721Uses tensorflow for automatic differentiation. This is also using raw HMC parametrized by
number of steps (see Pyro for commentary). This is the most verbose of the libraries
reviewed here, which can be good (more control) and bad (this was four lines in
PyMC3)I actually had a bug for a while where I had
w_dist = tfd.Normal(loc=tf.zeros(ndims), scale=1.0, name="w"). This defines a valid
model, because tfp is so flexible, but the wrong model.. The
computations are also all vectorized: I can sample 1,000 chains in about the same time I sample 4
chains,
which seems pretty unique.Blog post coming on
that. These are some reasons why one might write a high level, user friendly API on top of
this library and name it PyMC4.
import tensorflow as tf
import tensorflow_probability as tfp
tfd = tfp.distributions
X_tensor = tf.convert_to_tensor(X, dtype='float32')
@tf.function
def target_log_prob_fn(w):
w_dist = tfd.Normal(loc=tf.zeros((ndims, 1)), scale=1.0, name="w")
w_prob = tf.reduce_sum(w_dist.log_prob(w))
y_dist = tfd.Normal(loc=tf.matmul(X_tensor, w), scale=0.1, name="y")
y_prob = tf.reduce_sum(y_dist.log_prob(y_obs.reshape(-1, 1)))
return w_prob + y_prob
# Initialize the HMC transition kernel.
num_results = 1000
num_burnin_steps = 500
adaptive_hmc = tfp.mcmc.SimpleStepSizeAdaptation(
tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=target_log_prob_fn,
num_leapfrog_steps=4,
step_size=0.01),
num_adaptation_steps=int(num_burnin_steps * 0.8))
samples, is_accepted = tfp.mcmc.sample_chain(
num_results=num_results,
num_burnin_steps=num_burnin_steps,
current_state=tf.zeros((ndims, 1)),
kernel=adaptive_hmc,
trace_fn=lambda _, pkr: pkr.inner_results.is_accepted)
Edward 2
version 0.8.0-dev20190721Uses tensorflow for automatic differentiation. This is a submodule of TensorFlow Probability
that provides a flexible API based on the ideas from the popular Edward libraryThe same developer, so the similarity is not
surprising.. Note the biggest change from the tfp code above is that defining
the joint
probability is a bit cleaner, but we still use the same code to actually get the samples.
from tensorflow_probability import edward2 as ed
import tensorflow as tf
X_tensor = tf.convert_to_tensor(X, dtype='float32')
def linear_regression(X):
w = ed.Normal(loc=tf.zeros((ndims, 1)), scale=1.0, name="w")
y = ed.Normal(loc=tf.matmul(X, w), scale=0.1, name='y')
return y
log_joint = ed.make_log_joint_fn(linear_regression)
def target_log_prob_fn(w):
return log_joint(X_tensor, w=w, y=y_obs.reshape(-1, 1))
# Below here is from TFP section
num_results = 1000
num_burnin_steps = 500
adaptive_hmc = tfp.mcmc.SimpleStepSizeAdaptation(
tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=target_log_prob_fn,
num_leapfrog_steps=4,
step_size=0.01),
num_adaptation_steps=int(num_burnin_steps * 0.8))
ed_samples, is_accepted = tfp.mcmc.sample_chain(
num_results=num_results,
num_burnin_steps=num_burnin_steps,
current_state=tf.zeros((ndims, 1)),
kernel=adaptive_hmc,
trace_fn=lambda _, pkr: pkr.inner_results.is_accepted)
Numpyro
version 0.1.0Uses jax for automatic differentiation. Built by the same people who work on Pyro, and
includes a very cool iterative implemenation of the NUTS algorithmI have not looked closely enough to see if this is
"NUTS as in the 2013 paper", or "NUTS as in the algorithm implemented by Stan/PyMC3.". Note
the import section looks a little overwhelming, but jax is a wonderful library that wraps
numpy functionality. A more typical usage would be to import jax.numpy as np, so that no
one was any wiser that you are using autodiff. This looks very similar to the Pyro implementation, but
notice I am jumping through fewer hoops to convert my numpy arrays into torch tensors.
import jax.numpy as jnp
from jax import random
import numpyro.distributions as dist
from numpyro.handlers import sample
from numpyro.hmc_util import initialize_model
from numpyro.mcmc import mcmc
def model(X):
w = sample('w', dist.Normal(jnp.zeros(ndims), jnp.ones(ndims)))
y = sample('y', dist.Normal(jnp.matmul(X, w), 0.1 * jnp.ones(ndata)), obs=y_obs)
rng = random.PRNGKey(0)
init_params, potential_fn, constrain_fn = initialize_model(rng, model, X=X)
num_warmup, num_samples = 1000, 2000
# Run NUTS.
npyro_samples = mcmc(num_warmup, num_samples, init_params,
potential_fn=potential_fn,
trajectory_length=10,
constrain_fn=constrain_fn)
Brancher
version 0.3.4NOTE: Very new library. As far as I can tell, this does not implement MCMC (yet?), nor can I calculate log probabilities to verify my implementation. It looks nice, though!
Uses pytorch for automatic differentiation. This came out this week, and I gave it a test
drive, but could not do much damage with it yet.
from brancher.variables import ProbabilisticModel from brancher.standard_variables import NormalVariable from brancher import inference import brancher.functions as BF import torch X_tensor = torch.as_tensor(X, dtype=torch.float32) # Model w = NormalVariable(loc=torch.zeros(ndims), scale=1., name="w") y = NormalVariable(loc=BF.matmul(X_tensor, w), scale=0.1, name="y") y.observe(y_obs) model = ProbabilisticModel([w, y])
PyMC4
version 0.0.1NOTE: Still working on the API. Don't use this yet.
Uses tensorflow probability (and hence TensorFlow) for automatic differentiation. I am including this for what the model definition syntax is looking like right now, though some work needs to happen to wire the model through to the proper TensorFlow Probability functions.
import pymc4
import tensorflow as tf
@pymc4.model()
def linear_model():
w = yield pymc4.distributions.Normal('w', mu=np.zeros((5, 1)), sigma=1.)
y = yield pymc4.distributions.Normal('y', mu=tf.matmul(X, w), sigma=0.1)
PyProb
version 0.13.0Joins emcee in not using a gradient based sampler -- here we use importance sampling. I also
had to cheat a little bit and take 50,000 samples instead of 1,000 to get the estimates to be
reasonable.
The samples are pretty fast, but I suspect this does not scale to higher
dimensionsNow I have a paper to read about how importance sampling can
scale. (but I also suspect
that this does allow for defining a more broad class of models than most of the other languages).
Defining this model was pretty nice, and I am surprised more languages do not subclass a
Model
object.
import torch
import pyprob
from pyprob import Model
from pyprob.distributions import Normal
class LinearModel(Model):
def forward(self):
X_tensor = torch.as_tensor(X, dtype=torch.float32)
w = pyprob.sample(Normal(torch.zeros(ndims), torch.ones(ndims)))
y = Normal(torch.matmul(X_tensor, w), 0.1 * torch.ones(ndata))
pyprob.observe(y, name='y_obs')
return w
model = LinearModel()
posterior = model.posterior_distribution(
num_traces=50_000,
inference_engine=pyprob.InferenceEngine.IMPORTANCE_SAMPLING,
observe={'y_obs': torch.as_tensor(y_obs, dtype=torch.float32)})
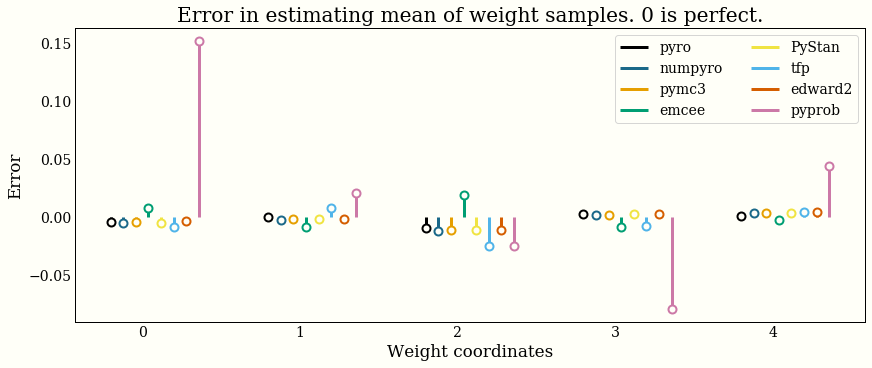
Accuracy
I have confidence that all the libraries implement their algorithm of choice correctly, and I tried to write a simple model that would provide some reasonable samples from the correct posterior. This section only tries to verify that my implementations are reasonable.
To say again with a stronger font, any errors shown here are (probably) MCMC error, and should not be interpreted as an implementation being "good" or "bad". It is a bigger, more careful job to benchmark performance, which is definitely not what this chart does.